Deguelin applied to chrysanthemum aphis
deguelin.RdQuantal assay data from an experiment where the insectide deguelin was applied to Macrosiphoniella sanborni.
Usage
data(deguelin)Format
A data frame with 6 observations on the following 4 variables.
dosea numeric vector of doses applied
log10dosea numeric vector of logarithm-transformed doses
ra numeric vector contained number of dead insects
na numeric vector contained the total number of insects
Details
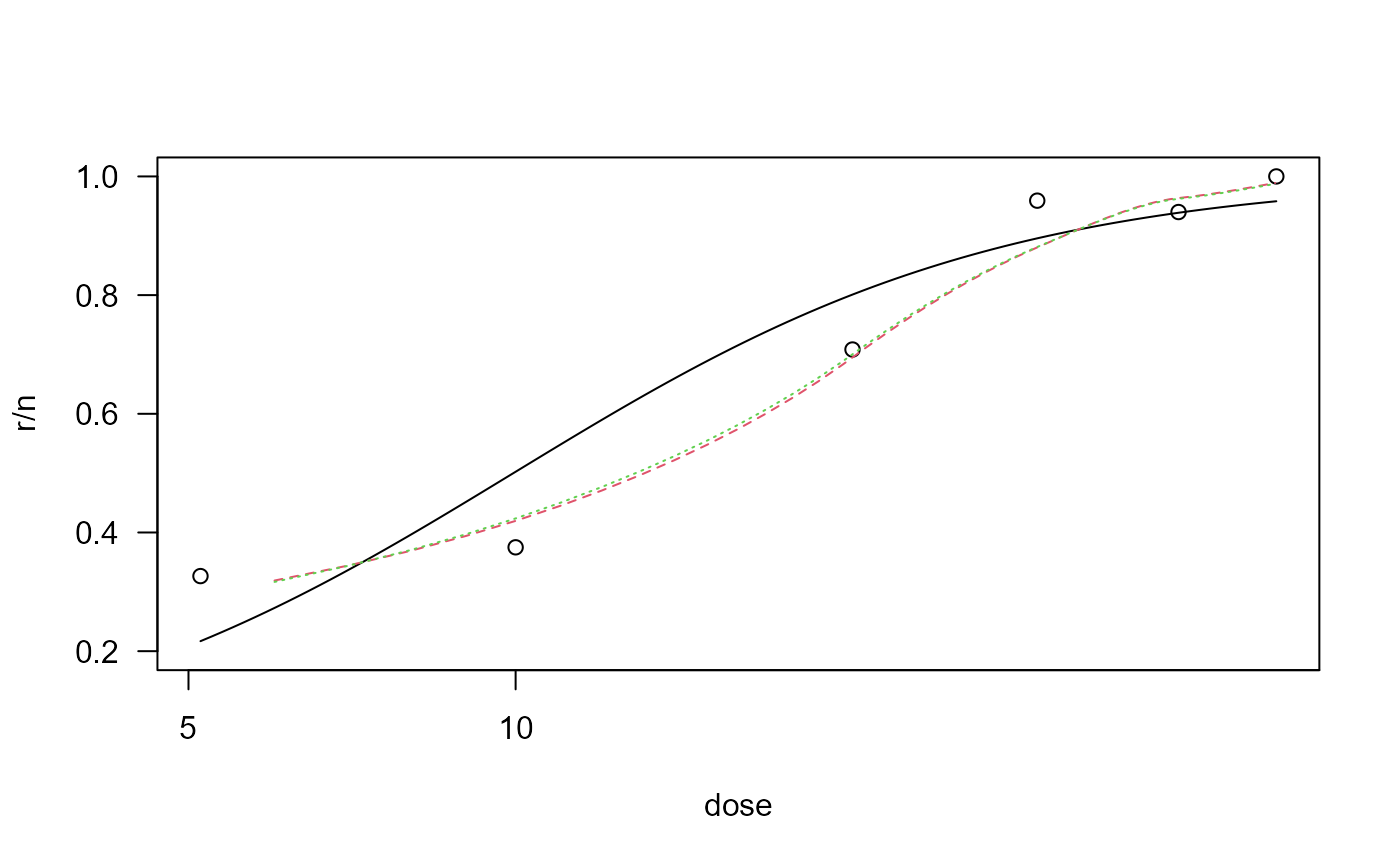
The log-logistic model provides an inadequate fit.
The dataset is used in Nottingham and Birch (2000) to illustrate a semiparametric approach to dose-response modelling.
Source
Morgan, B. J. T. (1992) Analysis of Quantal Response Data, London: Chapman & Hall/CRC (Table 3.9, p. 117).
References
Notttingham, Q. J. and Birch, J. B. (2000) A semiparametric approach to analysing dose-response data, Statist. Med., 19, 389–404.
Examples
library(drc)
## Log-logistic fit
deguelin.m1 <- drm(r/n~dose, weights=n, data=deguelin, fct=LL.2(), type="binomial")
modelFit(deguelin.m1)
#> Goodness-of-fit test
#>
#> Df Chisq value p value
#>
#> DRC model 4 13.375 0.0096
summary(deguelin.m1)
#>
#> Model fitted: Log-logistic (ED50 as parameter) with lower limit at 0 and upper limit at 1 (2 parms)
#>
#> Parameter estimates:
#>
#> Estimate Std. Error t-value p-value
#> b:(Intercept) -1.93709 0.22390 -8.6514 < 2.2e-16 ***
#> e:(Intercept) 9.95219 0.92186 10.7958 < 2.2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
## Loess fit
deguelin.m2 <- loess(r/n~dose, data=deguelin, degree=1)
## Plot of data with fits superimposed
plot(deguelin.m1, ylim=c(0.2,1))
lines(1:60, predict(deguelin.m2, newdata=data.frame(dose=1:60)), col = 2, lty = 2)
lines(1:60, 0.95*predict(deguelin.m2,
newdata=data.frame(dose=1:60))+0.05*predict(deguelin.m1, newdata=data.frame(dose=1:60), se = FALSE),
col = 3, lty=3)