Network Formation Assay Data
nfa.RdNeurotoxicity test using a network formation assay studying the inhibition of network formation at acrylamide exposure.
Usage
data(nfa)Format
A data frame with 45 observations on the following 4 variables.
chipchip ID
conc7 concentrations of acrylamide, ranging from 0-5mM
experimentfactor with levels 1 or 2 denoting two consecutive experiments
responseNumber of connections [%]
References
Frimat, JP, Sisnaiske, J, Subbiah, S, Menne, H, Godoy, P, Lampen, P, Leist, M, Franzke, J, Hengstler, JG, van Thriel, C, West, J. The network formation assay: a spatially standardized neurite outgrowth analytical display for neurotoxicity screening. Lab Chip 2010; 10:701-709.
Examples
data(nfa)
## Fit a four-parameter log-logistic model
nfa.m1 <- drm(response ~ conc, data = nfa, fct = LL.4())
summary(nfa.m1)
#>
#> Model fitted: Log-logistic (ED50 as parameter) (4 parms)
#>
#> Parameter estimates:
#>
#> Estimate Std. Error t-value p-value
#> b:(Intercept) 1.828165 0.429684 4.2547 0.0001184 ***
#> c:(Intercept) -3.879086 5.620773 -0.6901 0.4939982
#> d:(Intercept) 88.533870 2.366218 37.4158 < 2.2e-16 ***
#> e:(Intercept) 0.730916 0.086889 8.4121 1.813e-10 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error:
#>
#> 9.591243 (41 degrees of freedom)
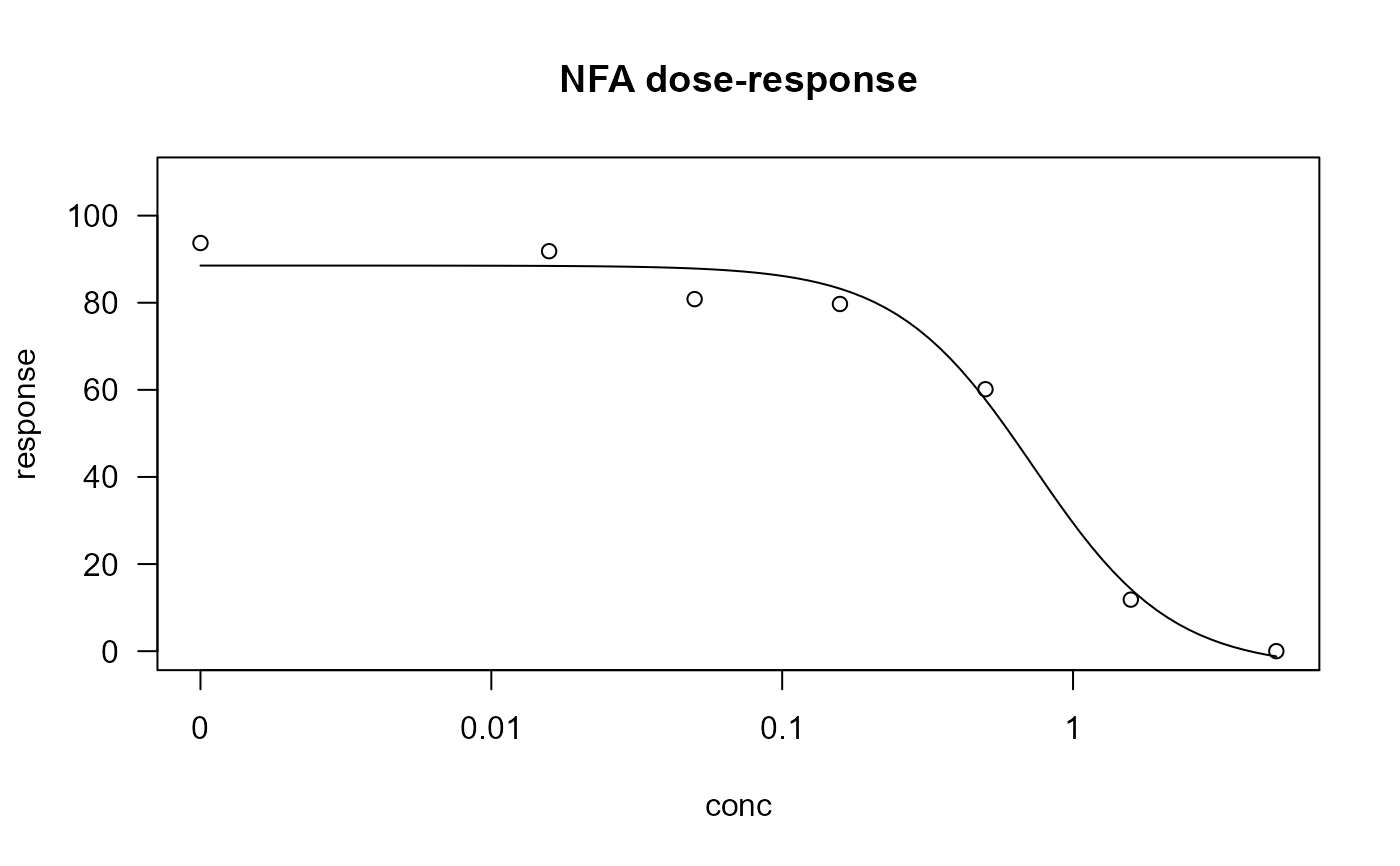
plot(nfa.m1, main = "NFA dose-response")